Practice, block vs. alternated degree-of-freedom ordering in 3D plate buckling¶
This notebook compares two strategies for positioning the degrees of freedom (DOFs) inside the global stiffness matrix and geometric stiffness matrix of a 3D plate buckling problem solved with the Ritz method using Legendre polynomial bases.
Both formulations solve the same physical problem — the only difference is the order in which the , , coefficients are stored in the global DOF vector .
| Strategy | Global DOF vector layout | Index of | Index of | Index of |
|---|---|---|---|---|
| Block | ||||
| Alternated |
where is the number of basis function triplets.
The two orderings are related by a permutation matrix such that , and consequently:
Since the eigenvalues of a generalized eigenvalue problem are invariant under simultaneous congruence transformation, both orderings must produce identical eigenvalues. This notebook verifies that claim and examines how DOF ordering affects sparsity patterns and solver behavior.
1Install required libraries¶
!python -m pip install buckling structsolve > tmp.txt2Import required libraries¶
import numpy as np
import matplotlib.pyplot as plt
import plotly.graph_objects as go
from scipy.special import roots_legendre
from scipy.sparse import csc_matrix
from structsolve import lb
from buckling.legendre import vecf, vecfxi3Approximation space and quadrature¶
We define the polynomial expansion orders () for the displacement field along the axes. The total degrees of freedom is , and the number of basis function triplets is .
Integration points and weights are computed using Gauss-Legendre quadrature. The number of integration points () is chosen to exactly integrate the stiffness terms formulated by the polynomial bases.
m1 = m2 = 7
m3 = 5
M = m1 * m2 * m3 # DOFs per displacement component
N = 3 * M # Total DOFs
print(f'm1 = {m1}, m2 = {m2}, m3 = {m3}')
print(f'DOFs per component (M) = {M}')
print(f'Total DOF (N) = {N}')
pts1, weights1 = roots_legendre(2*m1 - 1)
pts2, weights2 = roots_legendre(2*m2 - 1)
pts3, weights3 = roots_legendre(2*m3 - 1)m1 = 7, m2 = 7, m3 = 5
DOFs per component (M) = 245
Total DOF (N) = 735
4Material, geometry, and boundary conditions¶
Isotropic material properties and plate dimensions are defined. A fundamental compressive pre-stress state (sigxx = -1.0) is applied to evaluate the geometric stiffness.
Boundary constraints (xit1, xir1, etc.) are defined to enforce essential boundary conditions directly on the trial functions.
E = 200.0e9
nu = 0.3
G = E / (2 * (1 + nu))
a = 0.3
b = 0.1
h = 0.003
sigxx = -1.0
xit1, xir1 = 0, 1
xit2, xir2 = 0, 1
etat1, etar1 = 0, 1
etat2, etar2 = 0, 1
zetat1, zetar1 = 1, 1
zetat2, zetar2 = 1, 15Pre-allocation and helper functions¶
Both DOF orderings share the same set of shape-function vectors ( and their spatial derivatives), stiffness matrices and , and the same in-place outer-product helper addouter.
The only difference between the two approaches is where each coefficient is placed within these length- vectors — the physics (strain definitions, constitutive law, Jacobian) is identical.
def make_arrays():
"""Allocate fresh working arrays for one assembly pass."""
Su = np.zeros(N); Sv = np.zeros(N); Sw = np.zeros(N)
Sux = np.zeros(N); Svx = np.zeros(N); Swx = np.zeros(N)
Suy = np.zeros(N); Svy = np.zeros(N); Swy = np.zeros(N)
Suz = np.zeros(N); Svz = np.zeros(N); Swz = np.zeros(N)
buff = np.zeros((N, N))
K = np.zeros((N, N))
KG = np.zeros((N, N))
return (Su, Sv, Sw, Sux, Svx, Swx, Suy, Svy, Swy,
Suz, Svz, Swz, buff, K, KG)
def addouter(matrix, vec1, vec2, buff):
"""In-place addition of outer product to a target matrix."""
np.outer(vec1, vec2, out=buff)
matrix += buff5.1DOF ordering: block layout¶
In the block (sequential) ordering, all -coefficients come first, then all -coefficients, then all -coefficients:
In code, this maps to slice-based indexing:
Su[:M] = flat # u-component
Sv[M:2*M] = flat # v-component
Sw[2*M:] = flat # w-component63D numerical integration — block DOF assembly¶
The linear stiffness matrix and geometric stiffness matrix are built via numerical integration over the 3D domain using the block DOF layout.
(Su, Sv, Sw, Sux, Svx, Swx, Suy, Svy, Swy,
Suz, Svz, Swz, buff, K_block, KG_block) = make_arrays()
for xi, wxi in zip(pts1, weights1):
P_xi = vecf(m1, xi, xit1, xir1, xit2, xir2)
Px_xi = vecfxi(m1, xi, xit1, xir1, xit2, xir2)
for eta, weta in zip(pts2, weights2):
P_eta = vecf(m2, eta, etat1, etar1, etat2, etar2)
Px_eta = vecfxi(m2, eta, etat1, etar1, etat2, etar2)
for zeta, wzeta in zip(pts3, weights3):
P_zeta = vecf(m3, zeta, zetat1, zetar1, zetat2, zetar2)
Px_zeta = vecfxi(m3, zeta, zetat1, zetar1, zetat2, zetar2)
weight = wxi * weta * wzeta
# --- Block indexing: Su[:M], Sv[M:2M], Sw[2M:] ---
Pi, Pj, Pk = np.meshgrid(P_xi, P_eta, P_zeta, indexing='ij')
flat = (Pi*Pj*Pk).flatten()
Su[:M] = flat; Sv[M:2*M] = flat; Sw[2*M:] = flat
Pxi, Pj, Pk = np.meshgrid(Px_xi, P_eta, P_zeta, indexing='ij')
flat_x = (Pxi*Pj*Pk*(2/a)).flatten()
Sux[:M] = flat_x; Svx[M:2*M] = flat_x; Swx[2*M:] = flat_x
Pi, Pxj, Pk = np.meshgrid(P_xi, Px_eta, P_zeta, indexing='ij')
flat_y = (Pi*Pxj*Pk*(2/b)).flatten()
Suy[:M] = flat_y; Svy[M:2*M] = flat_y; Swy[2*M:] = flat_y
Pi, Pj, Pxk = np.meshgrid(P_xi, P_eta, Px_zeta, indexing='ij')
flat_z = (Pi*Pj*Pxk*(2/h)).flatten()
Suz[:M] = flat_z; Svz[M:2*M] = flat_z; Swz[2*M:] = flat_z
NL = 0
exx = Sux + NL*1/2*(Sux**2 + Svx**2 + Swx**2)
eyy = Svy + NL*1/2*(Suy**2 + Svy**2 + Swy**2)
ezz = Swz + NL*1/2*(Suz**2 + Svz**2 + Swz**2)
gxy = Suy + Svx + NL*(Sux*Suy + Svx*Svy + Swx*Swy)
gxz = Suz + Swx + NL*(Sux*Suz + Svx*Svz + Swx*Swz)
gyz = Svz + Swy + NL*(Suy*Suz + Svy*Svz + Swy*Swz)
C = E / ((1 - 2*nu)*(1 + nu))
Sxx = C * ((1 - nu)*exx + nu*eyy + nu*ezz)
Syy = C * (nu*exx + (1 - nu)*eyy + nu*ezz)
Szz = C * (nu*exx + nu*eyy + (1 - nu)*ezz)
Txy = G * gxy; Txz = G * gxz; Tyz = G * gyz
detJ = a * b * h / 8
addouter(K_block, detJ*weight*Sxx, exx, buff)
addouter(K_block, detJ*weight*Syy, eyy, buff)
addouter(K_block, detJ*weight*Szz, ezz, buff)
addouter(K_block, detJ*weight*Txy, gxy, buff)
addouter(K_block, detJ*weight*Txz, gxz, buff)
addouter(K_block, detJ*weight*Tyz, gyz, buff)
addouter(KG_block, detJ*weight*sigxx*Swx, Swx, buff)
print('Block assembly done.')Block assembly done.
6.1Eigenvalue solution — block DOF¶
eigvals_block, eigvecs_block = lb(csc_matrix(K_block), csc_matrix(KG_block),
sparse_solver=False)
eigvals_block_sparse, eigvecs_block_sparse = lb(csc_matrix(K_block), csc_matrix(KG_block),
sparse_solver=True)
D = E * h**3 / (12 * (1 - nu**2))
lbd_block = eigvals_block[0] * sigxx * b**2 * h / np.pi**2 / D
print(f'Block ordering — normalized buckling load: {lbd_block:.6f}')
print(f'First 5 eigenvalues: {eigvals_block[:5]}')Running linear buckling analysis...
Eigenvalue solver...
Removing null columns...
360 columns removed
finished!
eigh() solver...
finished!
finished!
first 5 eigenvalues:
1522040927.7001903
2282698902.6239214
7425108997.486525
8109202084.400536
14777273133.311518
Running linear buckling analysis...
Eigenvalue solver...
eigsh() solver (sigma=1.0)...
WARNING: Factor is exactly singular
aborted!
Removing null columns...
360 columns removed
finished!
eigsh() solver (sigma=4.1489519909504436e-10)...
finished!
finished!
first 5 eigenvalues:
1522040927.6879106
2282698902.658078
7425108997.522677
8109202084.386688
14777273133.059479
Block ordering — normalized buckling load: -9.355709
First 5 eigenvalues: [1.52204093e+09 2.28269890e+09 7.42510900e+09 8.10920208e+09
1.47772731e+10]
/opt/miniconda3/lib/python3.13/site-packages/structsolve/linear_buckling.py:93: RuntimeWarning: divide by zero encountered in divide
eigvals = -1./eigvals
/opt/miniconda3/lib/python3.13/site-packages/structsolve/linear_buckling.py:12: MatrixRankWarning: Matrix is exactly singular
y = spsolve(K, KG @ x)
7DOF ordering: alternated (interleaved) layout¶
In the alternated ordering, the three displacement components for each basis triplet are grouped together:
The -th basis triplet maps to global indices , , for , , respectively. In code, this maps to stride-based indexing:
Su[0::3] = flat # u-component (every 3rd entry starting at 0)
Sv[1::3] = flat # v-component (every 3rd entry starting at 1)
Sw[2::3] = flat # w-component (every 3rd entry starting at 2)This layout is common in finite-element codes where nodal DOFs are kept together for easier application of boundary conditions and contact constraints.
7.13D numerical integration — alternated DOF assembly¶
The identical triple Gauss-Legendre loop is executed, but the shape-function vectors are filled using the alternated (interleaved) index mapping. The physics (strain definitions, constitutive law, Jacobian) remain unchanged.
(Su, Sv, Sw, Sux, Svx, Swx, Suy, Svy, Swy,
Suz, Svz, Swz, buff, K_alt, KG_alt) = make_arrays()
for xi, wxi in zip(pts1, weights1):
P_xi = vecf(m1, xi, xit1, xir1, xit2, xir2)
Px_xi = vecfxi(m1, xi, xit1, xir1, xit2, xir2)
for eta, weta in zip(pts2, weights2):
P_eta = vecf(m2, eta, etat1, etar1, etat2, etar2)
Px_eta = vecfxi(m2, eta, etat1, etar1, etat2, etar2)
for zeta, wzeta in zip(pts3, weights3):
P_zeta = vecf(m3, zeta, zetat1, zetar1, zetat2, zetar2)
Px_zeta = vecfxi(m3, zeta, zetat1, zetar1, zetat2, zetar2)
weight = wxi * weta * wzeta
# --- Alternated indexing: Su[0::3], Sv[1::3], Sw[2::3] ---
Pi, Pj, Pk = np.meshgrid(P_xi, P_eta, P_zeta, indexing='ij')
flat = (Pi*Pj*Pk).flatten()
Su[0::3] = flat; Sv[1::3] = flat; Sw[2::3] = flat
Pxi, Pj, Pk = np.meshgrid(Px_xi, P_eta, P_zeta, indexing='ij')
flat_x = (Pxi*Pj*Pk*(2/a)).flatten()
Sux[0::3] = flat_x; Svx[1::3] = flat_x; Swx[2::3] = flat_x
Pi, Pxj, Pk = np.meshgrid(P_xi, Px_eta, P_zeta, indexing='ij')
flat_y = (Pi*Pxj*Pk*(2/b)).flatten()
Suy[0::3] = flat_y; Svy[1::3] = flat_y; Swy[2::3] = flat_y
Pi, Pj, Pxk = np.meshgrid(P_xi, P_eta, Px_zeta, indexing='ij')
flat_z = (Pi*Pj*Pxk*(2/h)).flatten()
Suz[0::3] = flat_z; Svz[1::3] = flat_z; Swz[2::3] = flat_z
NL = 0
exx = Sux + NL*1/2*(Sux**2 + Svx**2 + Swx**2)
eyy = Svy + NL*1/2*(Suy**2 + Svy**2 + Swy**2)
ezz = Swz + NL*1/2*(Suz**2 + Svz**2 + Swz**2)
gxy = Suy + Svx + NL*(Sux*Suy + Svx*Svy + Swx*Swy)
gxz = Suz + Swx + NL*(Sux*Suz + Svx*Svz + Swx*Swz)
gyz = Svz + Swy + NL*(Suy*Suz + Svy*Svz + Swy*Swz)
C = E / ((1 - 2*nu)*(1 + nu))
Sxx = C * ((1 - nu)*exx + nu*eyy + nu*ezz)
Syy = C * (nu*exx + (1 - nu)*eyy + nu*ezz)
Szz = C * (nu*exx + nu*eyy + (1 - nu)*ezz)
Txy = G * gxy; Txz = G * gxz; Tyz = G * gyz
detJ = a * b * h / 8
addouter(K_alt, detJ*weight*Sxx, exx, buff)
addouter(K_alt, detJ*weight*Syy, eyy, buff)
addouter(K_alt, detJ*weight*Szz, ezz, buff)
addouter(K_alt, detJ*weight*Txy, gxy, buff)
addouter(K_alt, detJ*weight*Txz, gxz, buff)
addouter(K_alt, detJ*weight*Tyz, gyz, buff)
addouter(KG_alt, detJ*weight*sigxx*Swx, Swx, buff)
print('Alternated assembly done.')Alternated assembly done.
7.2Eigenvalue solution — alternated DOF¶
eigvals_alt, eigvecs_alt = lb(csc_matrix(K_alt), csc_matrix(KG_alt),
sparse_solver=False)
eigvals_alt_sparse, eigvecs_alt_sparse = lb(csc_matrix(K_alt), csc_matrix(KG_alt),
sparse_solver=True)
lbd_alt = eigvals_alt[0] * sigxx * b**2 * h / np.pi**2 / D
print(f'Alternated ordering — normalized buckling load: {lbd_alt:.6f}')
print(f'First 5 eigenvalues: {eigvals_alt[:5]}')Running linear buckling analysis...
Eigenvalue solver...
Removing null columns...
360 columns removed
finished!
eigh() solver...
finished!
finished!
first 5 eigenvalues:
1522040927.695189
2282698902.708363
7425108997.673563
8109202084.373646
14777273133.24723
Running linear buckling analysis...
Eigenvalue solver...
eigsh() solver (sigma=1.0)...
WARNING: Factor is exactly singular
aborted!
Removing null columns...
360 columns removed
finished!
eigsh() solver (sigma=3.018426128182952e-10)...
finished!
finished!
first 5 eigenvalues:
1522040927.6726487
2282698902.651802
7425108997.464223
8109202084.340463
14777273133.24027
Alternated ordering — normalized buckling load: -9.355709
First 5 eigenvalues: [1.52204093e+09 2.28269890e+09 7.42510900e+09 8.10920208e+09
1.47772731e+10]
8Permutation equivalence verification¶
We now verify that and are related by the permutation that maps block indices to alternated indices:
If the two systems are truly equivalent, then element-wise.
# Build the permutation: perm[block_index] = alternated_index
perm = np.zeros(N, dtype=int)
for n in range(M):
perm[n] = 3*n # u_n
perm[M + n] = 3*n + 1 # v_n
perm[2*M + n] = 3*n + 2 # w_n
inv_perm = np.argsort(perm)
K_permuted = K_block[np.ix_(inv_perm, inv_perm)]
KG_permuted = KG_block[np.ix_(inv_perm, inv_perm)]
print(f'max|K_permuted - K_alt| = {np.max(np.abs(K_permuted - K_alt)):.2e}')
print(f'max|KG_permuted - KG_alt| = {np.max(np.abs(KG_permuted - KG_alt)):.2e}')
print()
print('The matrices are identical under permutation => mathematically equivalent systems.')max|K_permuted - K_alt| = 0.00e+00
max|KG_permuted - KG_alt| = 0.00e+00
The matrices are identical under permutation => mathematically equivalent systems.
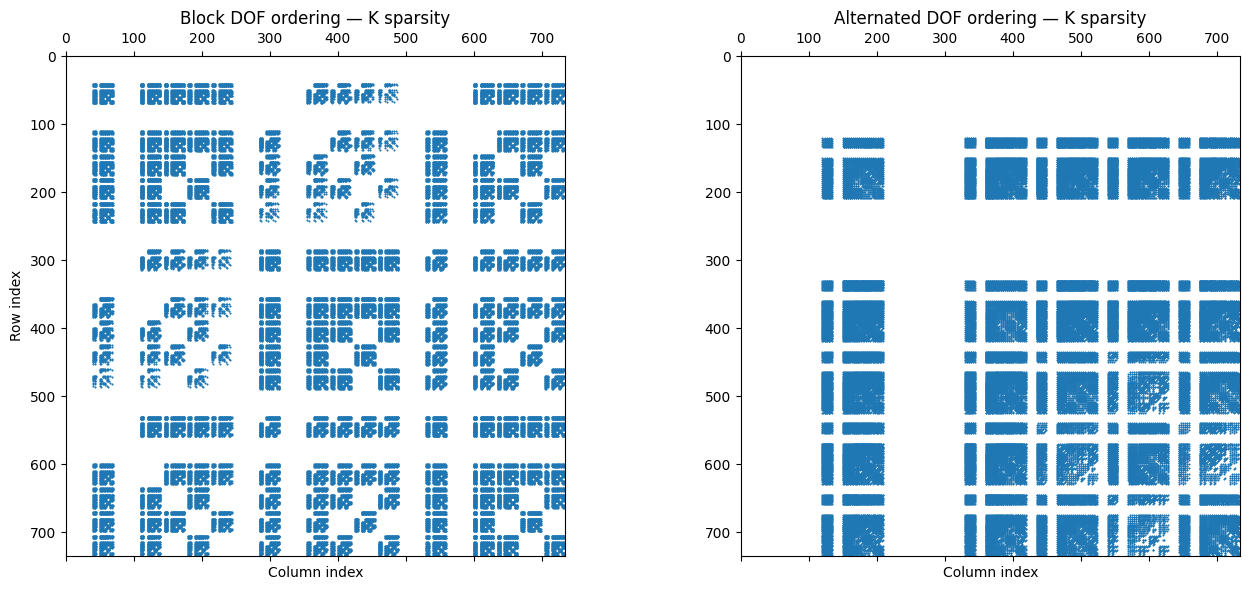
9Comparison of stiffness matrix sparsity patterns¶
Although the two systems have the same eigenvalues, their sparsity patterns differ dramatically. The block ordering produces a characteristic block structure with three dense diagonal blocks (, , ) and off-diagonal coupling blocks. The alternated ordering distributes the non-zeros more uniformly, producing a banded or semi-banded pattern.
These structural differences affect:
Memory layout: block ordering is more cache-friendly during assembly since consecutive DOFs often share the same basis function type.
Sparse storage: CSC/CSR compression ratios differ between the two patterns.
Iterative solvers: preconditioner effectiveness and convergence rates depend on the bandwidth and clustering of non-zeros.
fig, axes = plt.subplots(1, 2, figsize=(14, 6))
thr = 1e-6 * np.max(np.abs(K_block))
axes[0].spy(np.abs(K_block) > thr, markersize=0.5, aspect='equal')
axes[0].set_title('Block DOF ordering — K sparsity')
axes[0].set_xlabel('Column index')
axes[0].set_ylabel('Row index')
axes[1].spy(np.abs(K_alt) > thr, markersize=0.5, aspect='equal')
axes[1].set_title('Alternated DOF ordering — K sparsity')
axes[1].set_xlabel('Column index')
nnz_block = np.count_nonzero(np.abs(K_block) > thr)
nnz_alt = np.count_nonzero(np.abs(K_alt) > thr)
print(f'Non-zeros (block): {nnz_block} ({100*nnz_block/N**2:.1f}%)')
print(f'Non-zeros (alternated): {nnz_alt} ({100*nnz_alt/N**2:.1f}%)')
plt.tight_layout()
plt.show()
#fig.savefig('BasicPrinciples-DOF-ordering-sparsity.pdf', bbox_inches='tight')
#fig.savefig('BasicPrinciples-DOF-ordering-sparsity.jpg', bbox_inches='tight')
Non-zeros (block): 71915 (13.3%)
Non-zeros (alternated): 71915 (13.3%)
10Comparison of eigenvalues¶
The table below compares the sparse and dense solvers, showing that they produce identical eigenvalues. The observed differences are due to floating-point arithmetic in the assembly loop, on the order of machine precision.
n_compare = min(10, len(eigvals_block))
print(f'{"Mode":<6} {"Block (dense)":>18} {"Alt (dense)":>18} {"Block (sparse)":>18} {"Alt (sparse)":>18} {"Rel. diff":>12}')
print('-' * 94)
for i in range(n_compare):
rel = abs(eigvals_block[i] - eigvals_alt[i]) / abs(eigvals_block[i]) if eigvals_block[i] != 0 else 0
eb_sp = eigvals_block_sparse[i] if i < len(eigvals_block_sparse) else float('nan')
ea_sp = eigvals_alt_sparse[i] if i < len(eigvals_alt_sparse) else float('nan')
print(f'{i+1:<6} {eigvals_block[i]:>18.6e} {eigvals_alt[i]:>18.6e} {eb_sp:>18.6e} {ea_sp:>18.6e} {rel:>12.2e}')Mode Block (dense) Alt (dense) Block (sparse) Alt (sparse) Rel. diff
----------------------------------------------------------------------------------------------
1 1.522041e+09 1.522041e+09 1.522041e+09 1.522041e+09 3.29e-12
2 2.282699e+09 2.282699e+09 2.282699e+09 2.282699e+09 3.70e-11
3 7.425109e+09 7.425109e+09 7.425109e+09 7.425109e+09 2.52e-11
4 8.109202e+09 8.109202e+09 8.109202e+09 8.109202e+09 3.32e-12
5 1.477727e+10 1.477727e+10 1.477727e+10 1.477727e+10 4.35e-12
6 2.864875e+10 2.864875e+10 2.864875e+10 2.864875e+10 1.58e-12
7 3.956282e+10 3.956282e+10 3.956282e+10 3.956282e+10 6.17e-12
8 4.235453e+10 4.235453e+10 4.235453e+10 4.235453e+10 8.24e-12
9 4.476558e+10 4.476558e+10 4.476558e+10 4.476558e+10 3.55e-12
10 4.895995e+10 4.895995e+10 4.895995e+10 4.895995e+10 9.32e-11
113D visualization of buckling mode shapes¶
The eigenvector coefficients are extracted differently depending on the DOF ordering:
| DOF ordering | -coefficients | -coefficients | -coefficients |
|---|---|---|---|
| Block | eigvecs[0:M, mode] | eigvecs[M:2M, mode] | eigvecs[2M:3M, mode] |
| Alternated | eigvecs[0::3, mode] | eigvecs[1::3, mode] | eigvecs[2::3, mode] |
Once extracted, each coefficient vector is reshaped into a tensor and contracted with the 1D basis evaluation matrices via np.einsum to reconstruct the 3D displacement field.
The visualization below uses the block ordering eigenvectors.
num_modes = 3
scale = 100
nx, ny, nz = 40, 20, 3
x_1d = np.linspace(0, a, nx)
y_1d = np.linspace(0, b, ny)
z_1d = np.linspace(-h/2, h/2, nz)
xi_1d = 2 * x_1d / a - 1
eta_1d = 2 * y_1d / b - 1
zeta_1d = 2 * z_1d / h - 1
P_xi = np.array([vecf(m1, xi, xit1, xir1, xit2, xir2) for xi in xi_1d])
P_eta = np.array([vecf(m2, eta, etat1, etar1, etat2, etar2) for eta in eta_1d])
P_zeta = np.array([vecf(m3, zeta, zetat1, zetar1, zetat2, zetar2) for zeta in zeta_1d])
U_field = np.zeros((num_modes, nx, ny, nz))
V_field = np.zeros((num_modes, nx, ny, nz))
W_field = np.zeros((num_modes, nx, ny, nz))
for mode in range(num_modes):
# Block ordering: [u | v | w]
c_U = eigvecs_block[0:M, mode].reshape((m1, m2, m3))
c_V = eigvecs_block[M:2*M, mode].reshape((m1, m2, m3))
c_W = eigvecs_block[2*M:3*M, mode].reshape((m1, m2, m3))
U_field[mode] = np.einsum('ijk, xi, yj, zk -> xyz', c_U, P_xi, P_eta, P_zeta)
V_field[mode] = np.einsum('ijk, xi, yj, zk -> xyz', c_V, P_xi, P_eta, P_zeta)
W_field[mode] = np.einsum('ijk, xi, yj, zk -> xyz', c_W, P_xi, P_eta, P_zeta)
X, Y, Z = np.meshgrid(x_1d, y_1d, z_1d, indexing='ij')
fig = go.Figure()
for mode in range(num_modes):
X_def = X + scale * U_field[mode]
Y_def = Y + scale * V_field[mode]
Z_def = Z + scale * W_field[mode]
W_amp = W_field[mode]
lines_x, lines_y, lines_z, lines_w = [], [], [], []
for j in range(ny):
for k in range(nz):
lines_x.extend(X_def[:, j, k].tolist() + [None])
lines_y.extend(Y_def[:, j, k].tolist() + [None])
lines_z.extend(Z_def[:, j, k].tolist() + [None])
lines_w.extend(W_amp[:, j, k].tolist() + [0.0])
for i in range(nx):
for k in range(nz):
lines_x.extend(X_def[i, :, k].tolist() + [None])
lines_y.extend(Y_def[i, :, k].tolist() + [None])
lines_z.extend(Z_def[i, :, k].tolist() + [None])
lines_w.extend(W_amp[i, :, k].tolist() + [0.0])
for i in range(nx):
for j in range(ny):
lines_x.extend(X_def[i, j, :].tolist() + [None])
lines_y.extend(Y_def[i, j, :].tolist() + [None])
lines_z.extend(Z_def[i, j, :].tolist() + [None])
lines_w.extend(W_amp[i, j, :].tolist() + [0.0])
fig.add_trace(go.Scatter3d(
x=lines_x, y=lines_y, z=lines_z, mode='lines',
line=dict(color=lines_w, colorscale='Jet', width=3,
showscale=True, colorbar=dict(title=f'W (Mode {mode+1})', x=0.85)),
name=f'Mode {mode+1}', opacity=0.9, visible=(mode == 0)
))
buttons = []
for mode in range(num_modes):
vis = [False] * num_modes
vis[mode] = True
buttons.append(dict(label=f'Mode {mode+1}', method='update',
args=[{'visible': vis},
{'title': f'Buckling Mode Shape: Mode {mode+1}'}]))
fig.update_layout(
updatemenus=[dict(active=0, buttons=buttons, x=0.05, xanchor='left',
y=1.05, yanchor='top', direction='down', showactive=True)],
scene=dict(xaxis_title='X', yaxis_title='Y', zaxis_title='Z', aspectmode='data'),
title='Buckling Mode Shape: Mode 1', showlegend=False,
margin=dict(l=0, r=0, b=0, t=40)
)
fig.show()12Conclusion¶
Block and alternated DOF orderings produce stiffness matrices that are identical up to a permutation. The eigenvalue spectrum is therefore invariant — the physics does not depend on how array indices are assigned. As the comparison table above confirms, both the sparse (eigsh) and dense (eigh) solvers now produce identical eigenvalues for both orderings.
However, DOF ordering has important numerical consequences:
Sparsity pattern: block ordering yields a block-structured matrix; alternated ordering yields a banded pattern. These affect sparse storage efficiency, preconditioner design, and fill-in during factorization.
Sparse solver and the shift parameter:
structsolve.lbwithsparse_solver=Trueuseseigshin Cayley shift-invert mode, which requires a shift parametersigmaclose to the target eigenvalues. When is singular (as here), the solver first catches theRuntimeError: Factor is exactly singularand falls back toremove_null_colsbefore retrying. The_estimate_sigmafunction instructsolveestimates an appropriate shift via one inverse-iteration step (solving and computing the Rayleigh quotient), producing asigmaof order 10-10 — close to the actual eigenvalues — so thateigshconverges correctly regardless of DOF ordering.eighrobustness: the dense solvereighcomputes all eigenvalues via direct factorization without a shift parameter, giving correct and consistent results regardless of DOF ordering. It is the safer choice for small-to-moderate systems where the cost is acceptable.